U.S. Geological Survey Open-File Report 94-645
This report is preliminary and has not been reviewed for conformity with U.S. Geological Survey editorial standards (or with the North American Stratigraphic Code). Any use of trade, product, or firm names is for descriptive purposes only and does not imply endorsement by the U.S. Government.
Initially the empirical relationship between climate and components of the sediment was determined using a multiple regression technique (Imbrie and Kipp, 1971). In these studies sea-floor sediments were examined to determine the percentage of various species of planktonic foraminifera present in them. Supposing that the distribution of foraminiferal assemblages depended strongly on the extremes of annual sea-surface temperature (SST), the foraminiferal assemblages (refined through use of varimax factor analysis) were regressed against the average SST during the coolest and warmest months of the year. The result was a set of transfer functions, equations that could be used to estimate cool and warm SST from the faunal composition of a sediment sample. Assuming that the ecological preference of the species had remained constant throughout the last several hundred thousand years, these transfer functions could be used to estimate SSTs during much of the late Pleistocene.
Hutson (1980) and Overpeck, Webb, and Prentice (1985) proposed an alternative approach to estimating paleoclimatic parameters. Their "method of modern analogs" revolved not around the existence of a few climatically-sensitive faunal assemblages but rather on the expectation that similar climatic regimes should foster similar faunal and floral assemblages. From a large pool of modern samples, those few are selected whose faunal compositions are most similar to a given fossil sample. Paleoclimate estimates are derived using the climatic character of only the most similar modern samples, the modern analogs of the fossil sample.
This report describes how to use the program ANALOG to carry out the method of modern analogs. It is assumed that the user has faunal census estimates of one or more fossil samples, and one or more sets of faunal data from modern samples. Furthermore, the user must understand the taxonomic categories represented in the data sets, and be able to recognize taxa that are or may be considered equivalent in the analysis.
ANALOG provides the user with flexibility in input data format, output
data content, and choice of distance measure, and allows the user to
determine which taxa from each modern and fossil data file are compared.
Most of the memory required by the program is allocated dynamically, so
that, on systems that permit program segments to grow, the program
consumes only as many system resources as are needed to accomplish its
task.
System requirements
Blank lines in the run description file are ignored. Comments may be included by inserting the pound sign `#' anywhere on a line; the pound sign and all chararcters to the end of the line are considered part of the comment and are ignored by the program. Lines may begin with any number of spaces or tab characters to enhance the file's readability. Names and keywords can be in upper or lower case or a mixture, but under UNIX, file names must be given with the correct letter case, or the program won't find the files. File names should not contain braces (`{' or `}') since these indicate the grouping of items in the run description file.
The run description file consists of a sequence of keywords followed in some cases by file names or descriptive words. Items may be listed in any order, but the grouping described by braces below must be maintained. This keeps related information together. An example is shown in figure 1.
The following words are keywords, and cannot be used as the names of files:
basis
sample
data
transform
meta
distance
report
name
closest
verbose
To carry out a complete analysis, the run description file must contain at
least one basis specification indicating the modern data base and
one sample specification indicating the fossil samples. All
other statements in the run description file are optional.
The general form of the basis specification is
basis {
data file_name: format
transform file_name
meta file_name: format
}
The sample statement is identical but the keyword sample is
used in place of basis.To include a data set in the analysis, give its group (basis or sample) followed by an opening brace, and specify the following information:
The group is delimited using braces so the program will know which transformation file and which meta data file go with each basis file. Only the data file specification is actually required; the other statements are optional but will typically be needed.
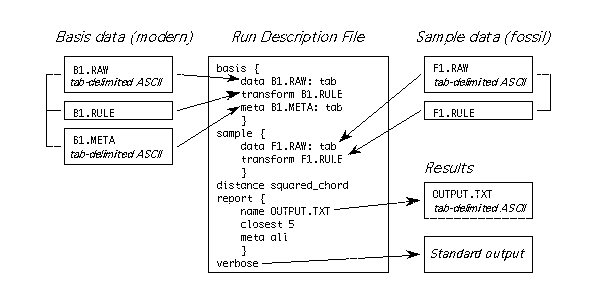
Figure 1. Information indicated in the run description file. The basis data files are shown as boxes on the left. The sample data files are shown on the right. Braces connect the raw, rule, and meta data files in the figure, but these are typically separate physical files on the disk. Results are output to the file OUTPUT.TXT, and the input data are echoed to the standard output device because verbose is specified in the run description file. This figure is available also as io.eps, an encapsulated PostScript file.
Example: To read the CLIMAP modern core-top data base as a basis, include the following text in the run description file:
basis {
data climap.raw: tab
transform climap.rule
meta climap.meta: tab
}
Example: To read the file mycore.dat as a fossil
sample, include the following text in the run description file:
sample {
data mycore.dat: tab
transform mycore.rule
}
where measure name is one of the following (the underscores must be included as shown):
manhattan
euclidean
squared_euclidean
canberra
squared_chord
squared_chisquared
dot_product
correlation
jaccard
sorensen
Mathematical descriptions of the distance and similarity measures are
included in a separate document.
report {
output file_name: format
closest number
meta variable_name_1
meta variable_name_2
...
meta variable_name_n
}
name is optional (if omitted, results are directed to the
standard output device); its syntax is the same as the data and
meta statements in input file specifications.closest should be followed by an integer; if closest is not indicated, the value 1 is used.
meta should be followed by the name of a meta data variable, in which case the statement can be repeated, or by all if you want all of the meta data variables added to the output for each matching modern sample. If the meta data variable's name contains any spaces, you must replace them with underscores in this statement.
Example: The following report statement causes ANALOG to create a file called analog.out for the results, to list the ten modern samples most similar to each fossil sample, and to report for each such modern sample the meta data variables present in the CLIMAP core-top data base.
report {
name analog.out: tab
closest 10
meta Latitude_degrees
meta Latitude_minutes
meta Longitude_degrees
meta Longitude_minutes
meta SST_cool
meta SST_warm
}
Since the transformation file allows you to rearrange the taxa found in one file so that they are commensurate with those in another, there will generally be one transformation file for every raw data file in the analysis.

Figure 2. Use of transformation files to make data from two different input files commensurate. Data file A has the variables A,B,C, and D, which are typically the names of biological taxa. Data file B has variables C, Da, Db, and E. In order to calculate distance coefficients properly, the variables must be the same in each data set. The user determines that taxa C, D, and E are present in both data sets, and notes that taxon E is composed of both types A and B, while taxon D is composed of types Da and Db. For data file A, the counts of taxa A and B are combined to calculate the counts of taxon E. A and B themselves are then ignored since these taxa are not tabulated separately in data file B. Likewise, data file A does not break taxon D into the two parts Da and Db, while data file B does. So the rules for data file B specify the creation of taxon D from Da and Db, and Da and Db are subsequently ignored. Finally, both data sets are transformed (after the addition and deletion of variables) by conversion to proportions (divide by sum). This figure is available also as an encapsulated PostScript file.
The transformation file is interpreted much like the run description file; blank lines are ignored, comments are indicated with the pound sign, spaces or tabs may precede keywords and taxon names, and keywords are not case sensitive. Certain words are recognized as keywords, and should not be used as taxon identifiers:
new
add
subtract
multiply
divide
ignore
all
by
The names of variables taken from the data file must be used here exactly
as they appear in the data file, including punctuation and letter case,
except that spaces embedded in the names must be replaced by
underscores. This is required because the code that parses the
transformation file uses spaces to separate words. By replacing spaces
with underscores, the different words of the taxon identifier are kept
together. The parser then converts underscores back into spaces in order
to compare names from the transformation file with names in the raw data
file. This also means that underscores should not appear in the names of
raw data variables.
Three directives can be given in transformation files: ignore,
new, and arithmetic operations.
ignore
ignore causes ANALOG to disregard the named variable. Its syntax
is ignore variable_name.
Examples:
ignore Depth_(mbsf)
ignore Indeterminates
new new_name {
add old_name_1
add old_name_2
...
add old_name_3
}
The new variable exists only while ANALOG is running. It cannot be
retained because the program is not intended as a data set editing tool
and is not programmed to save raw data on disk.Examples:
new dupac {
add N._dutertrei
add Neogloboquadrina_pachyderma
}
new Tsuga {
add Tsuga_canadensis? # note included '?'
add Tsuga_heterophylla # and '_' replaces ' '
add Tsuga_mertensiana
}
Note that including a taxon in a new category doesn't automatically cause
that taxon to be ignored. It may be useful to explicitly ignore
raw taxa that were used to create new categories.
It is not necessary to specify variables in any particular order. ANALOG
will sort them alphabetically and will only compare taxa whose names match
exactly. Even spaces before and after the names are considered when
comparing names.
arithmetic operations
ANALOG interprets the ignore and new instructions and
copies the raw data values (including those computed from new
instructions) into a separate array. At that point it can carry out
arithmetic operations on this new data array. The operations permitted
are indicated using these keywords:
multiply
divide
add
subtract
The operation word is followed by a decimal number or one of the words
count
sum
ssq
mean
var
sdev
count refers to the number of valid (i.e. not missing) data values in the sample.
sum refers to the sum of the data values for the sample.
ssq refers to the sum of squares of the data values for the sample.
mean refers to the arithmetic average of the data values for the sample.
var refers to the variance of the data values for the sample.
sdev refers to the standard deviation of the data values for the sample.
The variance and standard deviation here are n-weighted, meaning
that the variance is calculated as the sum of squared deviations from the
mean divided by the number of values n, not n - 1. The standard
deviation is calculated as the square root of the variance.multiply and divide may be followed by the word by; the by will be ignored. The number of arithmetic operations is not limited.
Example:
To obtain percentages from counts, use the statements
divide by sum
multiply by 100
In practice, it is not necessary to multiply by 100, as proportions are
more appropriate for calculating distance measures than are percentages.
Input data formats
Three input file formats are currently supported: tab-delimited ASCII, the
ASCII format of the North American Pollen Database, and the strict format
of the CLIMAP Atlantic Ocean data set as described in the first SPECMAP
data release. Other input file formats may be added, but that requires
amending and recompiling the source code. The default input file format
is tab-delimited ASCII; to read an input file in any other format, the
format must be indicated in the run description file.
Tab-delimited ASCII
Tab-delimited ASCII indicates that the data are stored in a plain text
file, one sample per line, with variable values separated by the tab
character (ASCII 8). This format is typically generated when a
spreadsheet program is asked to save data as "Text Only". ANALOG
requires also that
Sample ID\tG. ruber\tG. sacculifer\tN. pachyderma KNR110/43PC 0-2cm\t100\t50\t0 P1AR92B8 0-2cm\t0\t0\t200
Usage: ANALOG run_description_file_name
You neglected to enter the name of the run description file on the command line.Error: could not open ...
The name given is misspelled or the file is unavailable. If the file is the run description, which controls subsequent execution, the program does not continue. If the file is a raw or meta data file, the program tries to continue without that file's data.Error: unexpected end of run description file
The run description file is incomplete or there is an opening brace that has no corresponding closing brace.Error: expected (string), got (string) ...
The run description or transformation file contains a syntax error. This message appears when one of the following rules is broken:Error: unexpected (string) ...
- File name must follow data keyword.
- Format description must follow colon after file name if colon is present.
- File name must follow transform keyword.
- File name must follow meta keyword in basis or sample groups.
- File name must follow name keyword in report group.
- Meta variable name or keyword all must follow meta keyword in report group.
- Opening brace must follow basis, sample, or report keywords.
- Taxon name must follow ignore, new, or add in transformation file.
- Opening brace must follow taxon name in new statement of transformation file.
- Name following distance keyword must be a recognized distance measure.
- Number must follow closest keyword in report group.
- Number or keyword sum must follow arithmetic operator in transformation file; keyword by may appear between them.
Extraneous text appears where a keyword or closing brace should be. Check to see if you misspelled a keyword.Warning: extra data at end of line ...
Warning: extra meta data at end of line ...
ANALOG counts the number of variables in a tab-delimited file by looking at the column headings. If there are fewer column headings than there are columns with data in them, the program notices the extra characters at the end of the line and informs the user that they exist. It then proceeds, ignoring the extra characters.Warning: found meta data but no numerical data for sample (name) ...
Warning: no meta data found for sample (name) ...
Warning: found temperature data but no numerical data for sample (name) ...
A sample ID for meta data must match exactly the ID for the same sample in the raw data file. If they do not, the program assumes that the two records refer to different samples. If the program encounters meta data for a sample for which it has no raw data or, after having read meta data, finds samples for which no meta data could be found, these warnings are issued.Error: card out of order ...
Error: data card (number) refers to wrong core ...
The SPECMAP data is quite specific about the order of lines referring to an individual core. Similarly, the core identifier must be the same on the data cards as on the master card.Error: expected NAPD ASCII header line, got (string) ...
Error: expected two numbers, got (string) ...
Warning: expected sample meta data, got (string) ...
NAPD ASCII format files must abide by specific guidelines for the arrangment of data. If these guidelines are not followed, ANALOG will be unable to continue. Meta data can be read only from the modern files (those whose extension is .m50). An attempt to read meta data from a fossil NAPD data file (extension .f70) will generate the warning indicating that the meta data could not be read.Error: could not enlarge ...
Error: could not allocate ...
An attempt to allocate memory dynamically has failed. This indicates that there is not enough memory to read, interpret, or analyze the data. On some systems it may be possible to increase the amount of memory that is available to the program.Error: new variable (name) already exists in data file (name)
Error: You asked to create AND ignore new variable (name) ...
Warning: new variable (name) refers to old variable (name), which does not occur ...
Warning: rule file (name) says to ignore (name), which isn't in (name)
These messages indicate discrepancies between the rule file and the data file. Where possible, ANALOG will continue, issuing a warning. Where it cannot proceed without additional information from the user, ANALOG issues an error message and stops.Warning: data from (name) contains variable (name) not found in (name)
If, after transformation, the sample data contains a taxon that is not present in the basis data, or vice versa, that taxon is ignored when computing the distance measure. A warning is issued because the condition may give misleading results even though the distance measures can still be calculated.Warning: could not create output file (name); using stdout instead.
The named file could not be created. Instead, results will be directed to the standard output device.
It is possible to compile the Macintosh version so that it will run under Macintosh System 6. As distributed, System 7 is required because the program makes a call to PBCatSearch, a toolbox routine that locates a file by name on a Macintosh volume. If you replace the reference to Mac_fopen with the standard C fopen in the file input.c, and do not include the input file macfiles.c in the project, the program will run under System 6, but the data files must then reside in the same folder as the program.
An ancillary program called
transpos is available for MS-DOS and UNIX systems. This
program transposes the rows and columns of a tab-delimited text file. A
typical use might be to get a list of column headings from a tab-delimited
text file. Cut the first line from the tab-delimited text file, then run
the program transpos, redirecting output to a disk file. The
output will contain a list of column headings, one per line.
Additional file formats
Additional input file formats may be implemented. The code to read the
new format must be supplied by the user, compiled, and linked with the
rest of the source code. To enable ANALOG to recognize the new file
format, it is necessary to add an array element to the static array
read_function in file input.c. The first member of the array
element is a character string that will be used to identify the format;
this string will be given as the format in the run description file. The
second member is a pointer to the function that reads raw data from this
type of file. The third member is a pointer to the function that reads
meta data from this type of file.
The actions taken by the new read functions may be complex. Briefly, the read function is given a pointer to a data base structure, whose members include the information specified in the run description file. If p is the pointer passed to the read function, then raw data are read from p->raw.filespec; the variable names are stored in p->raw.name_buffer; pointers to the names are stored in p->raw.name, and the number of raw variables is stored in p->raw.count. Raw data for each sample are stored in the arrays p->sample[n].raw. The meta data must be read likewise from p->meta.filespec, the number of variables in p->meta.count, the names in p->meta.name_buffer, and pointers to the names in p->meta.name. Meta data values are stored as strings in the array p->sample[n].meta_buffer, and pointers to those strings are stored in p->sample[n].meta. See the code in input.c for details.
Adding another output format is essentially the same, but is less complex,
because the output data are simpler in form. See the code in
results.c for clues.
References
Peter N. Schweitzer
Mail Stop 918, National Center
U.S. Geological Survey
Reston, VA 20192
Tel: (703) 648-6533
FAX: (703) 648-6560
email: pschweitzer@usgs.gov
Maintained by Peter Schweitzer