Fact Sheet 2012–3027
The Molecular Epidemiology of Aquatic Pathogens (MEAP)-IHNV DatabaseThe MEAP-IHNV database provides access to detailed data for anyone interested in IHNV molecular epidemiology, such as fish health professionals, fish culture facility managers, and academic researchers. The flexible search capabilities enable the user to generate various output formats, including tables and maps, which should assist users in developing and testing hypotheses about how IHNV moves across landscapes and changes over time. The MEAP-IHNV database is available online at http://gis.nacse.org/ihnv/ (fig. 1). The database contains records that provide background information and genetic sequencing data for more than 1,000 individual field isolates of the fish virus Infectious hematopoietic necrosis virus (IHNV), and is updated approximately annually. It focuses on IHNV isolates collected throughout western North America from 1966 to the present. The database also includes a small number of IHNV isolates from Eastern Russia. By engaging the expertise of the broader community of colleagues interested in IHNV, our goal is to enhance the overall understanding of IHNV epidemiology, including defining sources of disease outbreaks and viral emergence events, identifying virus traffic patterns and potential reservoirs, and understanding how human management of salmonid fish culture affects disease. Ultimately, this knowledge can be used to develop new strategies to reduce the effect of IHN disease in cultured and wild fish. 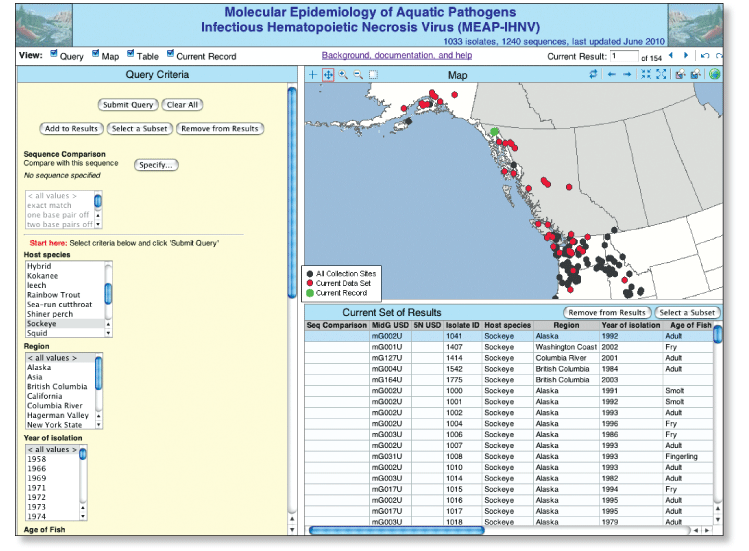
Figure 1. Screen view of database available at http://gis.nacse.org/ihnv. North American IHN VirusIHN virus is a well known pathogen of salmonid fish, with a host range including many species of salmon and trout (Bootland and Leong, 1999). Taxonomically it is the type species of the genus Novirhabdovirus, within the virus family Rhabdoviridae, which includes other well-known animal viruses, such as rabies virus and vesicular stomatitis virus. IHNV originated in western North America, where it is currently endemic to most Pacific watersheds that contain salmonid fish. In the 1970s and 1980s, IHNV was introduced to Europe and Asia by transport of virus-contaminated fish eggs. IHNV can be transmitted either horizontally through waterborne virus, or vertically as eggassociated virus. Infection in fish can result in lethal necrosis of the hematopoietic tissues of the kidney and spleen, and disease outbreaks can cause as much as 90 percent mortality depending on various host, virus, and environmental factors (fig. 2). IHNV is a significant impediment to rainbow trout farm culture and to hatchery and net-pen programs that grow salmon and trout. Due to the economic impact of IHNV, spawning adult and juvenile fish in most cultured salmonid populations in western North America are routinely surveyed for IHNV. Several molecular epidemiology studies of North American IHNV field isolates have revealed three major genetic subgroups of IHNV, designated U, M, and L because they occur in the upper, middle, and lower parts of the IHNV geographic range in North America (fig. 3; Emmenegger and others, 2000; Troyer and others, 2000; Emmenegger and Kurath, 2002; Garver and others, 2003; Kurath and others, 2003; Troyer and others, 2003; Rudakova and others, 2007). 
Figure 2. Signs of IHNV disease in rainbow trout (Oncorhynchus mykiss) include hemorrhage and exophthalmia (pop-eye) (photograph at left), and skin darkening relative to lighter colored healthy fish (photograph at right). 
Figure 3. Geographic ranges of the major subgroups of IHNV in western North America. Phylogenetic analyses have shown that U genogroup IHNV occurs in salmonid fish from Alaska, British Columbia, coastal Washington, and the Columbia River basin that extends throughout much of Washington, Oregon, and Idaho. IHNV in the M genogroup also occurs in the Columbia River basin, and recently has been detected in Washington coastal watersheds. IHNV in the L genogroup occurs in California and on the southern coast of Oregon. Although each genogroup can infect multiple fish host species, the U, M, and L groups show some host specificity for sockeye salmon (Oncorhynchus nerka), rainbow and steelhead trout (O. mykiss), and Chinook salmon (O. tshawytscha), respectively. IHNV cannot infect humans or other mammals. Genetic Typing Methods and Identification of GenotypesIHNV has a single-stranded, negative sense RNA genome of approximately 11,000 bases, which contains 6 genes. The G gene is approximately 1,610 nucleotides (nt) long and encodes the only protein on the outer surface of the enveloped virus particles. Standard genetic typing was done by determining the sequence of a variable 303 nt region in the middle of the G gene, referred to as the “midG” (fig. 4). Each different midG sequence is referred to as a “genotype”, and any virus isolates with the same midG sequence would be grouped within that genotype. Thus, some genotypes represent single isolates, and others may represent a large number of isolates. As described more thoroughly in the background section of the online database, each specific sequence genotype is referred to by a Universal Sequence Designator, or USD. For midG sequences the USDs are written in the format “mG###g.” For example, mG001U is the USD for a common midG sequence (genotype) in the U genogroup, and mG007M is the USD for a common genotype in the M genogroup. For some virus isolates, additional sequences have been determined, such as a partial N gene region, or complete G or N gene sequences. The database contains all sequence data available for each virus isolate. 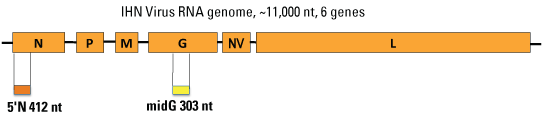
Figure 4. Regions of the IHN viral genome used for genetic typing. Each IHN virus particle contains a molecule of RNA that has the six viral genes, in the order shown. The G gene codes for the viral glycoprotein that covers the surface of the virus particles. Comparisons of genetic sequences of G genes from different IHNV isolates revealed a variable region in the middle of the G gene, termed the “midG,” that has been used extensively for IHNV genetic typing. An alternative region also used for typing is at the beginning of the viral N gene, which codes for the nucleocapsid protein that wraps up the viral RNA inside each virus particle. Origin of the Database InformationThe methods used for genetic typing of IHNV field isolates were developed in the mid-1990s at the U.S. Geological Survey (USGS), Western Fisheries Research Center (WFRC) in Seattle, Washington. Virus isolates and their associated background information were obtained from numerous state, Federal, and tribal fish health colleagues. A virus “isolate” is defined as the virus produced in cultured fish cells inoculated with a fish tissue sample collected from a specific fish, or pool of fish, at a specific time and location. Although most of the sequences in this database were generated at WFRC, collaborators at the Oregon Department of Fish and Wildlife, the University of California, Davis in California, and the Kamchatka Research Institute of Fishery and Oceanography in Russia, also contributed sequence data. How to Use IHNV Genotype Data for Epidemiological InsightsThe database can be searched by an actual genetic sequence, or by using the USD designation, to identify all isolates with identical genotypes. The USD system is helpful for working with the genetic typing data. Isolates with the same USD are likely to be more closely related than isolates with different USDs, although it is important to remember that additional sequence differences may occur outside the midG region. Isolates with the same USD may indicate epidemiological links that should be considered to reveal routes of virus transmission. When isolates have different USDs, the sequence differences serve as tags or markers that can be useful for tracking the virus over landscapes or through time. Single nt differences suggest close relationships, and may represent relatively recent divergence by mutation compared with isolates that have multiple nt differences. As a simple example of interpretation: two IHNV isolates from the same collection location differing in midG sequence by 1 nt could have arisen by a relatively recent mutation occurring at that location to create a variant, or by independent introductions of the two different types. However, two isolates from the same location differing by 5 nt in the midG more strongly suggests two introduction events. It is important to keep in mind that without additional testing, sequence differences at this level cannot be assumed to indicate any biological difference between isolates. However, the differences can be very useful for assessing virus traffic patterns and epidemiology. The database can be used to address localized epidemiology questions, such as identifying possible sources of disease outbreaks. As an example, if IHN disease occurs at a salmon hatchery, the genetic type of the virus would be determined from fish that died in the outbreak, and compared with the virus types in other potentially relevant samples, such as fish upstream in the water source, or fish from a farm that recently transported fish into the hatchery. If the virus from transported fish has several genetic differences from the dying fish, but the virus from fish in the water source has the same genotype, then it is more likely that the virus from infected fish in the hatchery water source caused the outbreak. Identification of possible routes of virus introduction can assist managers in deciding what actions to take to break chains of virus transmission. The database also can be used to look for larger patterns and associations for IHNV. For example, in the database 98 percent of the IHNV isolates from sockeye salmon were in the U genogroup, and 86 percent of the isolates from steelhead and rainbow trout were in the M genogroup. This pattern suggested differences in host-specificity between the major genogroups of IHNV that have been confirmed in laboratory studies. This understanding has greatly improved our ability to explain the ecology of IHNV in North America. For Additional InformationThe background section of the on-line database provides more detail on exactly what the database is and what it is not, as well as how to use it. This database does not represent all IHNV that has been isolated during fish health surveillance in North America, and the isolates included here do not necessarily represent consistent geographic or temporal coverage of the virus in North America. A more thorough explanation of development of the database is available in Emmenegger and others (2011). References CitedBootland, L.M., and Leong, J.C., 1999, Infectious hematopoietic necrosis virus, in Woo, P.T.K., and Bruno, D.W., eds., Fish diseases and disorders: New York, CAB International, v. 3, p. 57-121. Emmenegger, E.J., Kentop, E., Thompson, T.M., Pittam, S., Ryan, A., Keon, D., Carlino, J.A., Ranson, J., Life, R.B., Troyer, R.M., Garver, K.A., and Kurath, G., 2011, Development of an aquatic pathogen database (AquaPathogen X) and its utilization in tracking emerging fish virus pathogens in North America: Journal of Fish Diseases, no. 34, p. 579–587. Emmenegger, E.J., and Kurath, G., 2002, Genetic characterization of infectious hematopoietic necrosis virus of coastal salmonid stocks in Washington State: Journal of Aquatic Animal Health, no. 14, p. 25–34. Emmenegger, E.J., Meyer, T.R., Burton, T.O., and Kurath, G., 2000, Genetic diversity and epidemiology of infectious hematopoietic necrosis virus in Alaska: Diseases of Aquatic Organisms, no. 40, p. 163–176. Garver, K.A., Troyer, R.M., and Kurath, G., 2003, Two distinct phylogenetic clades of IHN virus overlap within the Columbia River basin: Diseases of Aquatic Organisms, no. 55, p. 187–203. Kurath, G., Garver, K.A., Troyer, R.M., Emmenegger, E.J., Einer-Jensen, K., and Anderson, E.D., 2003, Phylogeography of infectious hematopoietic necrosis virus in North America: Journal General Virology, no. 84, p. 803–814 (DOI 10.1099/ vir.0.18771-0). Rudakova, S.L., Kurath, G., and Bochkova, E.V., 2007, Occurrence and genetic typing of infectious hematopoietic necrosis virus in Kamchatka Russia: Diseases of Aquatic Organisms, no. 75, p. 1–11. Troyer, R.M., and Kurath, G., 2003, Molecular epidemiology of infectious hematopoietic necrosis virus reveals complex virus traffic and evolution within southern Idaho aquaculture: Diseases of Aquatic Organisms, no. 55, p. 175–185. Troyer, R.M., LaPatra, S., and Kurath, G., 2000, Genetic analyses reveal unusually high diversity of infectious hematopoietic necrosis virus in rainbow trout aquaculture: Journal of General Virology, no. 81, p. 2,823–2,832. |
First posted March 19, 2012 For additional information contact: Part or all of this report is presented in Portable Document Format (PDF); the latest version of Adobe Reader or similar software is required to view it. Download the latest version of Adobe Reader, free of charge. |
Kurath, Gael, 2012, An online database for IHN virus in Pacific Salmonid fish—MEAP-IHNV: U.S. Geological Survey Fact Sheet 2012–3027, 4 p.